
In November 2022, Sid Sijbrandij — the CEO of GitLab, a man who'd built a 2,500-person company without a single office — was told he had a six-centimeter mass growing from his T5 vertebra. Osteosarcoma. A rare bone cancer that mostly strikes teenagers.
He was 45 and otherwise healthy.
Two years of chemotherapy, radiation, and surgery left him battered but in remission. Then, in 2024, the cancer came back. Standard options were exhausted. His oncologist suggested finding a clinical trial.
Sid did not find a clinical trial. He went founder mode.
Today, his cancer is undetectable. His immune system was rewired — tumor-infiltrating T cells went from 19% to 89%. And every piece of data from his treatment — 25 terabytes of genomic, transcriptomic, and imaging data — is publicly available on Google Cloud.
This is the story of how he did it, why the tools are all open-source, and how we packaged the entire framework into an agent skill you can install right now.
Manager Mode vs. Founder Mode Medicine
Standard oncology works like manager-mode management. The physician knows best. Treatments are tried sequentially — if A fails, try B. Diagnostics are ordered conservatively. The patient complies.
This works for most conditions. It fails catastrophically when:
- Standard treatments are exhausted
- The cancer is rare (limited clinical trial data)
- The patient has the capability and motivation to do more
Sid flipped the model. He recruited Jacob Stern — a director at 10x Genomics — as the de facto CEO of his care. He built clinical and scientific advisory boards. He wrote a health document that grew past 1,000 pages.
And he discovered something surprising about the regulatory landscape: the FDA was fast. Every Individual Patient Expanded Access application (Form 3926) was approved within 48 hours. The real bottleneck was hospital IRBs — institutional review boards where a single member could block access to experimental treatment.
The system wasn't broken at the top. It was broken in the middle.
The Three Pillars
Sid's approach reduces to three principles that generalize beyond his specific cancer.
Pillar 1: Maximal Diagnostics
Standard clinical panels — a gene panel here, a CT scan there — give you a summary. Sid wanted the full picture.
He ran everything: whole genome sequencing from five providers, whole exome sequencing, bulk RNA-seq, single-cell RNA-seq (tumor biopsies and serial blood draws), three independent liquid biopsy platforms (Signatera, Northstar, Personalis), organoid drug testing, mass response drug testing, and PET scans with novel tracers most oncologists have never ordered.
The key diagnostic move: he insisted on cryopreserved tissue alongside the standard formalin-fixed samples. Formalin preserves cell morphology but destroys RNA. Without frozen tissue, the single-cell sequencing that changed everything would have been impossible.
Pillar 2: Personalized Therapeutic Development
Rather than waiting for clinical trials, Sid's team designed treatments based on his specific tumor biology. Five versions of neoantigen vaccines — peptide and mRNA — targeting mutations unique to his cancer. Oncolytic viruses engineered to kill tumor cells and release antigens. NK cell therapy. Gene therapy. Radioligand therapy accessed in Germany.
The regulatory mechanism: FDA Form 3926. One page. Approved in 48 hours. The FDA is not the enemy.
Pillar 3: Parallel Treatment
Standard oncology: try one thing, wait three months, scan, repeat. Sid's approach: run compatible therapies simultaneously and use liquid biopsy to measure what's working in real time.
During peak treatment, he was simultaneously receiving:
- Checkpoint inhibitors (Dostarlimab + Ipilimumab) — remove immune brakes
- Neoantigen vaccines (JLF + CeGaT peptides) — train immune recognition
- Oncolytic virus (AdaPT-001) — kill tumor cells, release antigens
- NK cell therapy (SNK-01) — innate immune killing
- Radioligand therapy (225Ac-FAPi) — targeted radiation to FAP+ cells
Each modality attacks from a different angle. The combination turned a "cold" tumor hot.
The Breakthrough Nobody Expected
Here's where it gets interesting for anyone who works with data.
Standard genomic panels — the kind you get from Tempus or Foundation Medicine — showed nothing actionable beyond the known driver mutations. The tumor was, by conventional metrics, immunologically boring.
But the single-cell RNA sequencing told a different story.
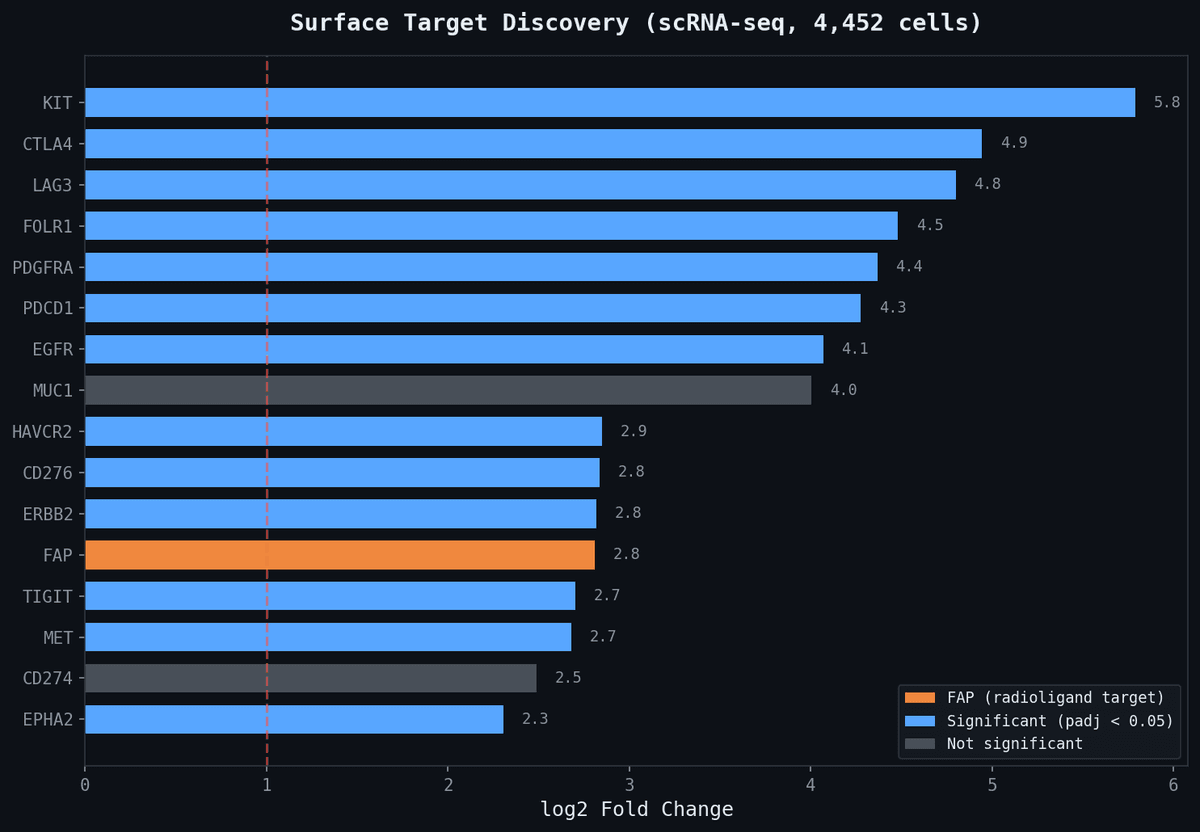
When Sid's team clustered the tumor cells by gene expression, they found massive overexpression of fibroblast activation protein (FAP) — a surface protein involved in wound healing that tumors hijack to build protective stroma. This isn't a standard biomarker for osteosarcoma. No gene panel tests for it. No WES report highlights it.
It took research-grade transcriptomics to see it.
And FAP is targetable.
Step 1: A 68Ga-FAP PET scan — gallium-68 conjugated to a FAP-targeting ligand — confirmed the tumor lit up. The target was validated in vivo.
Step 2: The same targeting molecule, now carrying a therapeutic payload — first lutetium-177 (beta emitter), then actinium-225 (alpha emitter) — delivered radiation directly to FAP-expressing cells.
Step 3: The tumor shrank enough for a second surgery at Memorial Sloan Kettering.
Step 4: Post-surgical tissue analysis showed the immune transformation — 19% to 89% T cell infiltration. The parallel immunotherapy had converted an immune desert into an immune battlefield.
The diagnostic insight that enabled all of this — FAP overexpression — was invisible to standard clinical tools.
The Pipeline Is Open Source
Every bioinformatics tool used in Sid's approach is available on GitHub:
| Step | Tool | Stars |
|---|---|---|
| DNA alignment | BWA | — |
| Variant calling | GATK Mutect2 | — |
| Copy number analysis | ASCAT | — |
| scRNA-seq analysis | Scanpy / Seurat | 2,000+ / 2,500+ |
| Neoantigen prediction | pVACtools | 300+ |
| MHC binding prediction | MHCflurry | 200+ |
| Vaccine ranking | Vaxrank (OpenVax) | 90 |
| End-to-end vaccine pipeline | openvax/neoantigen-vaccine-pipeline | 90 |
| mRNA vaccine synthesis guide | philfung/openvaxx | 67 |
The OpenVaxx project estimates a per-patient cost of $4,200 for in-house mRNA vaccine materials. The full neoantigen vaccine pipeline — from biopsy to ranked peptide candidates — runs on a 16-core machine with 64GB RAM in about 24-48 hours.
Sid went further and open-sourced his own data: 25 terabytes of WGS, WES, RNA-seq, scRNA-seq, long-read sequencing, spatial transcriptomics, and imaging — all publicly readable on Google Cloud via osteosarc.com.
Extending with AlphaFold and Protein Design
The sequence-based pipeline (pVACseq → MHCflurry → Vaxrank) predicts neoantigen binding from amino acid sequence alone. It's fast but approximate. We can do better.
AlphaFold Multimer can predict the 3D structure of a peptide-MHC complex — revealing whether the neoantigen physically fits in the MHC groove and which residues face outward for T cell recognition. Candidates that score high on both sequence-based affinity and structural confidence (ipTM > 0.5, PAE < 10) are stronger vaccine targets.
RFdiffusion (2,800 stars) + ProteinMPNN (1,700 stars) enable de novo therapeutic protein design. When no existing drug targets the discovered surface protein, you can design one from scratch: generate backbone geometries complementary to the target epitope, design amino acid sequences for each backbone, validate with ESMFold or AlphaFold2.
For radioligand therapy specifically, structural modeling of the FAP-ligand interface (AlphaFold DB: AF-Q12884-F1) can optimize binding and validate that diagnostic (68Ga) and therapeutic (177Lu/225Ac) payloads maintain equivalent affinity.
This is the structural biology layer that Sid's approach didn't use — but the next patient's should.
The Systemic Barriers Are Real
Despite having extraordinary resources, Sid encountered friction at every level:
Tissue access: Hospitals default to formalin-fixed tissue that destroys the RNA needed for transcriptomics. Getting cryopreserved samples requires advance coordination with both the surgeon and pathology department.
Data portability: Clinical sequencing companies provide summary reports. Getting raw FASTQ/BAM files requires formal data release requests that take 2-4 weeks.
IRB vetocracies: Hospital institutional review boards proved more obstructive than the FDA. No standardized timelines, no standardized criteria, and single members can block treatment access.
Economic misalignment: It costs $4.4 billion to develop and approve a new oncology drug. It costs $1 million to dose a single person with personalized therapy. The entire pharmaceutical industry is optimized for the former.
Sid launched Even One Ventures to fund companies attacking these barriers — seven portfolio companies working on scaling personalized oncology.
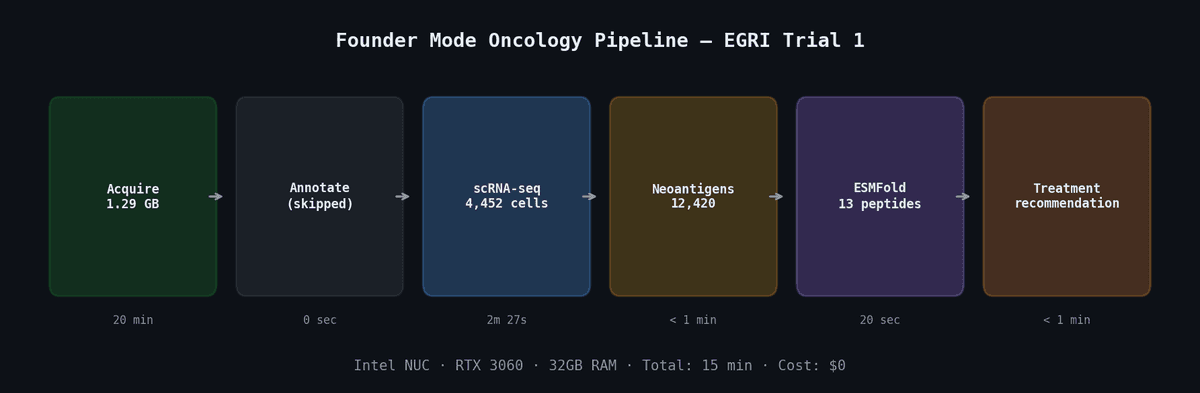
What We Built
We compiled all of this research into a structured agent skill:
npx skills add broomva/founder-mode-oncology
The founder-mode-oncology skill packages the entire framework — three pillars, decision workflow, open-source pipeline, treatment categories, regulatory navigation, MRD monitoring interpretation, and AlphaFold integration — into 7 files (1,321 lines) that work across 37+ AI agents.
It includes six reference documents:
- Diagnostics Pipeline — BWA → GATK → ASCAT → Scanpy → pVACtools, with hardware requirements
- Treatment Categories — 7 modalities with combination rationale and contraindications
- Regulatory Access — FDA Form 3926 walkthrough, IRB navigation, tissue access strategies
- MRD Monitoring — Three liquid biopsy platforms with interpretation guide and actionable thresholds
- Open-Source Tools — 25+ GitHub repos with descriptions and use cases
- Structural Biology — AlphaFold, RFdiffusion, ProteinMPNN workflows with quality thresholds
The research repo is also public: github.com/broomva/founder-mode-cancer
The Future Is Already Here
Elliot Hershberg, writing in Century of Bio, imagines a near future where early cancer detection triggers enrollment in a concierge oncology platform. AI agents analyze your medical history, order targeted diagnostics, generate bioinformatics reports, and recommend personalized combinations — checkpoint inhibitors (approaching generic pricing), off-the-shelf neoantigen vaccines (mRNA platform), and custom radioligand therapy. Total estimated cost: $175,000.
That's less than the current average cost of treating pancreatic cancer.
The technologies exist. Single-cell sequencing is commercially available. Neoantigen prediction runs on open-source software. AlphaFold is free. The FDA approves expanded access in 48 hours. mRNA vaccine platforms are proven.
The gap isn't technology. It's access, coordination, and willingness to operate outside the standard-of-care paradigm.
"It costs $1B to get a drug approved. But it costs $1M to dose a single person with personalized therapy." — Sid Sijbrandij
Sid is, in his own words, "the Kool-Aid Man breaking through the wall." He's living thirty years ahead. The question is how fast the rest of the system catches up.
Explore the data: osteosarc.com — 25TB of open genomic data
Install the skill: npx skills add broomva/founder-mode-oncology
Read the full case: Going Founder Mode on Cancer by Elliot Hershberg
Research repo: github.com/broomva/founder-mode-cancer